xGen™ RNA Library Preparation Kit
Accelerate your RNA-Seq transcriptomics research
The xGen RNA Library Prep Kit offers a fast NGS transcriptomics research workflow. Assemble libraries by either manual or automated workflow directly from 1st strand cDNA synthesis while using your preferred indexing options for the experiment.
xGen NGS—made for RNA library preparation.
Ordering
- Save time and cost—results in as little as 3.5 hours. Combined with the lower cost, you can process more samples in less time to increase your data output.
- Supports many workflows with indexing options, automation, and Normalase™ Technology—combinatorial dual indexing up to 96-plex, unique dual indexing up to 1536-plex, and enzymatic library normalization available; automation-friendly.
- Reliable functionality—maintains ligation efficiency at all supported inputs and no adapter titration required.
Product details
The xGen RNA Library Preparation Kit offers a fast, low-cost transcriptome sequencing workflow for Illumina® platforms. Our RNA library prep kit enables stranded RNA library construction directly from 1st strand cDNA without the requirement for 2nd strand cDNA synthesis and degradation, or template-switching methods. This easily automatable kit is compatible with upstream and downstream enrichment and depletion methods and supports a variety of indexing options. These include upstream mRNA enrichment and rRNA depletion modules as well as downstream xGen Hybridization capture workflows for targeted RNA sequencing.
Table 1. Research applications
| Application | xGen RNA Library Prep Kit |
|---|---|
| Gene expression profiling | ✓ |
| Research on disease-associated transcripts | ✓ |
| Total RNA + hybridization capture enrichment | ✓ |
Automation
The xGen RNA Library Prep Kit protocol is readily automatable. A 10% overage volume of reagents is supplied to accommodate automation, and additional reagent overage volume is available upon request.
Table 2. Product specifications
| Feature | Specification | Benefit |
|---|---|---|
| Input quantity | 100 ng to 1 μg total RNA 5 ng to 100 ng mRNA | Supports a wide input range |
| RNA types supported | Poly(A)-enriched mRNA Ribo-depleted RNA Total RNA | Supports most RNA applications |
| Technology | Adaptase™ tailing and ligation to 1st strand cDNA | Maintains strandedness (≥97%) No 2nd strand cDNA No adapter titration |
| Workflow time* | 3.5 hours | Less hands-on time |
| Kit reaction sizes | 16, 96, and 4x96 | Evaluation and adoption |
| Components provided | Fragmentation module RT module Library prep Polymerase | Complete solution for processing total, enriched, or depleted RNA from transcript to library |
| Indexing options | Combinatorial dual Unique dual Normalase | Flexible to sequencers, workflows, and applications |
| Multiplexing capability | Up to 1536 libraries | Save sequencing costs |
| Automation | Compatible with liquid handlers Custom packaging available | Adopt at scale |
*Workflow time is based upon incubation times and expected times for hands-on-steps. Actual workflow time may vary depending on individual factors in your laboratory.
Workflow
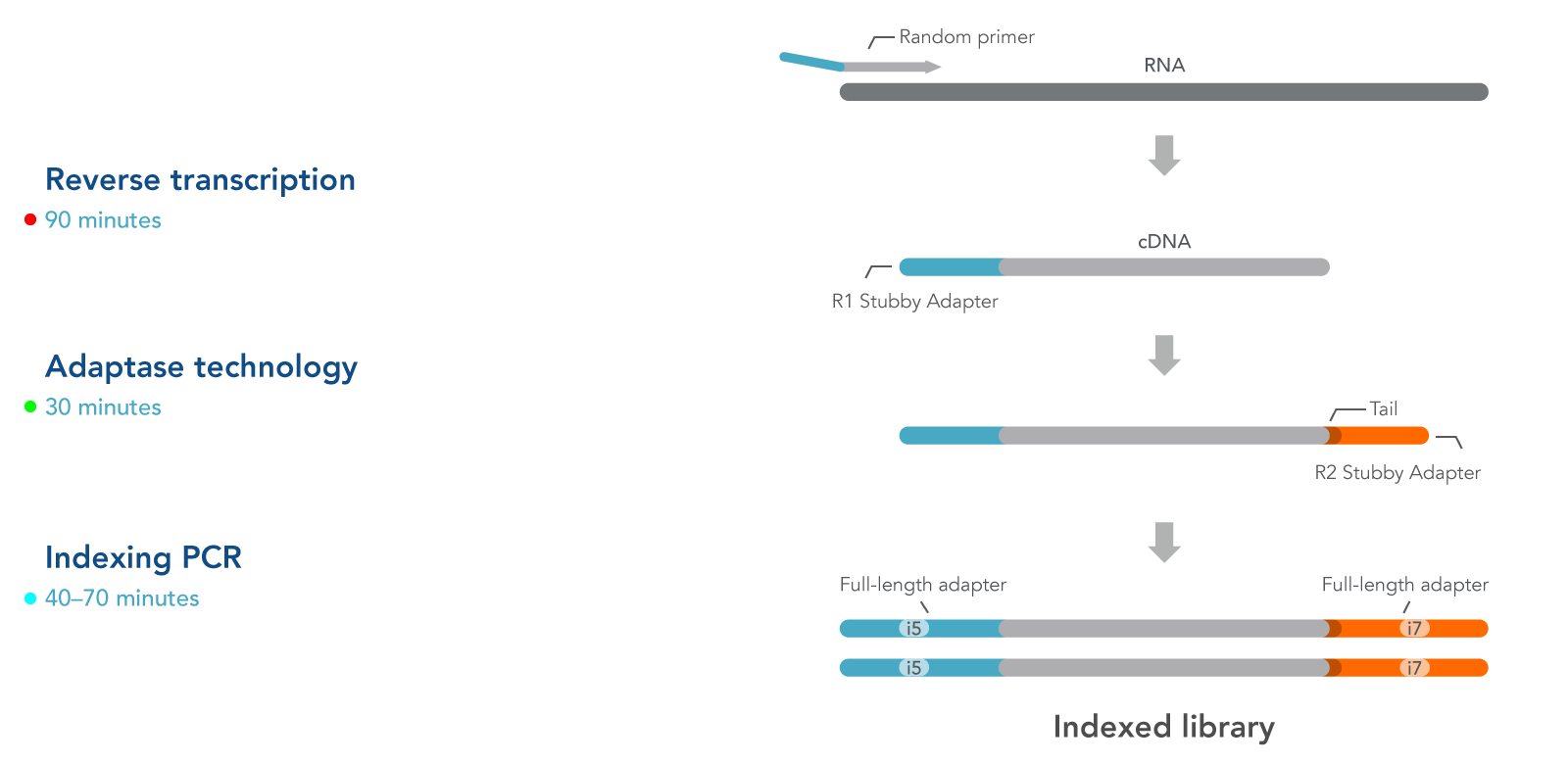
Reverse transcription—During this step, RNA fragmentation, reverse transcription, and incorporation of the R1 Stubby Adapter are performed, followed by a purification step.
Adaptase technology—This step performs 3’ tailing and ligation of the R2 Stubby Adapter to first strand cDNA
Indexing PCR—This step completes adapter sequences, adds sample-specific index sequences, and increases library yield, followed by a purification step.
Figure 1. xGen RNA Library Prep Kit workflow converts RNA directly into an indexed library. The workflow for creating an RNA-seq library using the xGen RNA Library Prep Kit includes a proprietary Adaptase technology step that adds a single-stranded R2 Stubby Adapter onto the 3’ end of the 1st strand cDNA product. Indexing primers are then used to amplify the library and add the full-length adapter sequence. See text for details.
Index and kit options
The xGen RNA Library Prep Kit uses primers that anneal to the Stubby Adapters added during the random priming and Adaptase steps, to be fully compatible with Illumina® sequencers. IDT supplies a variety of index configurations and strategies, including:
- Combinatorial dual indexing, up to 96 combinations
- Unique dual indexing, up to 1536 unique dual indices
Product data
Save time with direct conversion of 1st strand cDNA
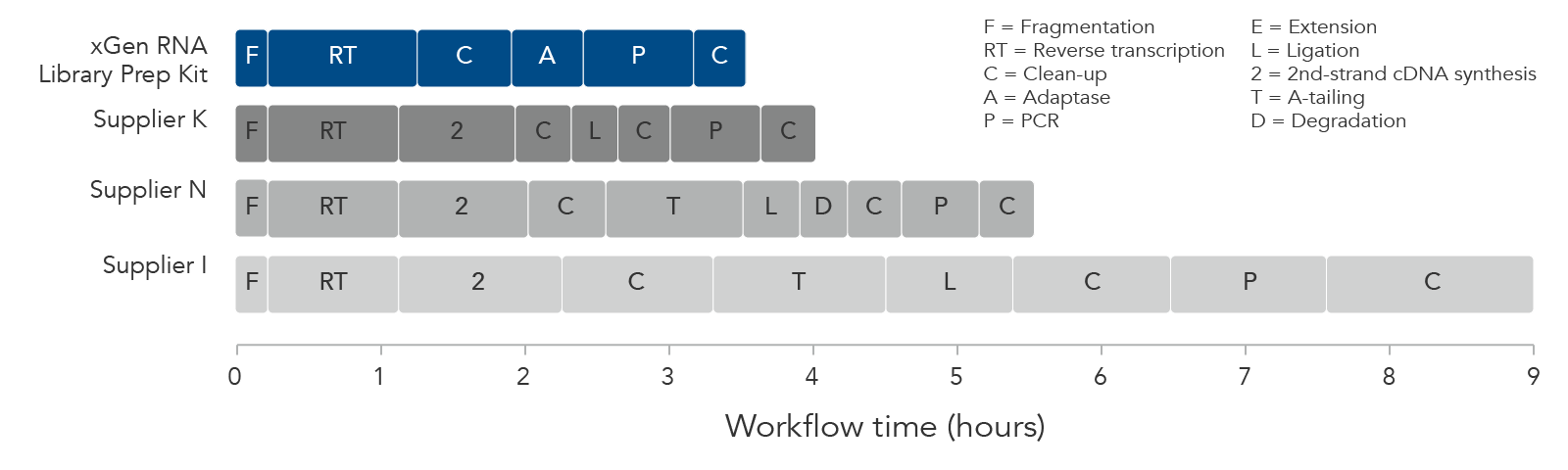
Incubation times for enzymatic and purification steps are shown according to vendor supplied user instructions.
Figure 2. Comparison of different workflow times for RNA-seq library preparation kits. The xGen RNA Library Prep Kit constructs library molecules directly from 1st strand cDNA synthesis, which reduces the time from fragmentation (F) to clean-up (C).
Reliable mapping and strandedness
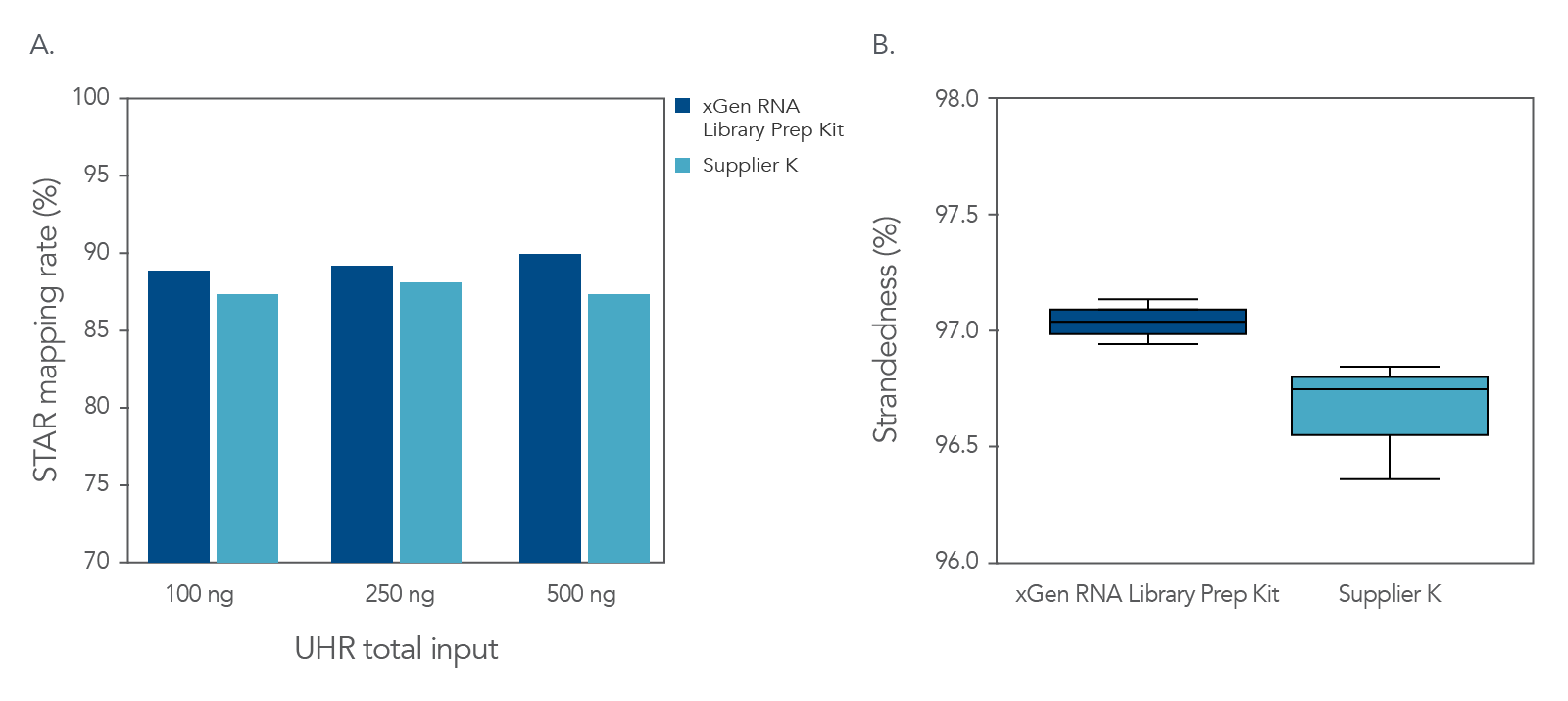
The xGen RNA Library Kit shows high mapping rates and maintenance of strandedness.
Figure 3. Reliable mapping rate and strandedness with the xGen RNA Library Prep Kit. (A) Universal Human Reference (UHR) Total RNA (Agilent) was enriched using the NEBNext Poly(A) mRNA Magnetic Isolation Module (NEB). Libraries were constructed from 50, 100, and 250 ng of enriched mRNA using the xGen RNA Library Prep Kit and the Supplier K’s kit according to the manufacturer’s instructions. (B) Human Brain mRNA (Clontech) was used to construct the xGen RNA and the Supplier K’s libraries from 1, 5, 10, 50, 100 ng of Human mRNA. The percent strandedness was determined by the average of the five input quantities. For both graph A and graph B, libraries were sequenced on a MiniSeqTM with 2 X 75 bp paired-end reads. Fastq files were downsampled to 3.8 million reads before analysis using STAR (mapping rate) or RNA-SeQC (strandedness).
Resources
Frequently asked questions
What are the new names of the Swift products?
The Swift products were rebranded and now belong to the xGen™ NGS product line. The following lists the old Swift product name, and then the new name hyperlinked to its current product page.
| Swift product name | New IDT product name |
|---|---|
| Accel-NGS® Adaptase® Module for Single-Cell Methyl-Seq | xGen Adaptase™ Module |
| Normalase® Amplicon Panels (SNAP) Core | xGen Amplicon Core |
| SARS-CoV-2 Additional Genome Coverage Panel | xGen SARS-CoV-2 Expanded Amplicon Panel |
| SARS-CoV-2 S Gene Panel | xGen SARS-CoV-2 SGene Amplicon Panel |
| SNAP Set 1A Combinatorial Dual Indexing Primers | xGen Amplicon CDI Primers |
| Swift Combinatorial Dual Indexing Primers | xGen CDI Primers |
| Swift Normalase® Combinatorial Dual Indexing Primers | xGen Normalase CDI Primers |
| Accel-NGS 1S Plus DNA Library | xGen ssDNA & Low-input DNA Library Prep |
| Swift 2S Sonic DNA Library | xGen DNA Library Prep MC |
| Swift 2S Sonic Flexible DNA Library | xGen DNA Library Prep MC UNI |
| Swift 2S Turbo v2 | xGen DNA Library Prep EZ |
| Swift 2S Turbo Flexible v2 DNA Library | xGen DNA Library Prep EZ UNI |
| Accel-NGS Methyl-Seq DNA Library | xGen Methyl-Seq Library Prep |
| Swift Normalase | xGen Normalase Module |
| Swift Rapid RNA Library | xGen RNA Library Prep Kit |
| Swift RNA Library | xGen Broad-Range RNA Library Prep |
Does the xGen™ RNA Library Prep Kit include upstream poly(A) enrichment or ribodepletion modules?
1. Total RNA
2. mRNA that has already been enriched by poly(A) selection
3. RNA that has already been depleted of rRNA
The xGen RNA Library Prep Kit begins with an RNA fragmentation and reverse transcription step. Therefore, if mRNA enrichment or ribodepletion is desired, one of the kits below is required and recommended for pre-processing. These kits have been tested for compatibility with the xGen RNA Library Prep Kit.
Note: View this xGen RNA Library Prep Kit Protocol for instructions on matching the output of these upstream modules with the initial steps of the xGen RNA Library Prep Kit.
| Method | Recommended kit | Alternate kit |
|---|---|---|
| Poly(A) mRNA enrichment | NEBNext® Poly (A) mRNA Magnetic Isolation Module Cat. No. E7490 |
Lexogen Poly (A) RNA Selection Kit Cat. No. 039.100 |
| Ribodepletion | Lexogen RiboCop rRNA Depletion Kit V1.2 (Human/Mouse/Rat) Cat. No. 037.24/96 |
NEBNext rRNA Depletion Kit (Human/Mouse/Rat) Cat. No. E6310S/L/X |
What indexing primers can I use with this xGen™ RNA Library Prep Kit?
The xGen™ RNA Library Prep Kits have an Indexing PCR step to complete the fully indexed adapter sequences following ligation of stubby adapters during the library prep workflows so they are fully compatible with Illumina® sequencers.
IDT supplies a variety of index configurations and strategies, including dual indexing as either:
- Combinatorial dual indexing, up to 96 combinations
- Unique dual indexing, up to 1536 combinations
What are the main steps for library construction using the xGen™ RNA Library Prep Kit?
The xGen™ RNA Library Prep Kit uses three main steps:
- Fragmentation & Reverse Transcription, where RNA is fragmented and converted to cDNA. Because the random primer is tailed, the 1st strand cDNA already contains the Read 1 Stubby Adapter.
- Adaptase, where the Read 2 Stubby Adapter is added to the 3’ end of 1st strand cDNA.
- Indexing PCR, where simultaneously the fully indexed adapter sequences are added, and the library is amplified to generate sufficient yield for sequencing.
What is the shelf life and recommended storage conditions for the xGen™ RNA Library Prep Kit?
The shelf life of the xGen RNA Library Prep Kit is at least 6 months from the time the product is delivered when stored as required at –20°C and handled according to the protocol.
The enzymes provided in this kit are temperature sensitive, and appropriate care should be taken during handling and storage. Upon receipt, store the kits at –20°C.
What are the input requirements for the xGen™ RNA Library Prep Kit?
How do I quantify and characterize xGen™ RNA libraries?
Following the recommended PCR cycles in the xGen RNA Library Prep Kit Protocol will result in a library concentration of at least 4 nM.
The xGen™ RNA Library Prep Kit can also be used with the Normalase™ Module to generate 2 nM or 4 nM normalized library pools.
Is titration of adapters required for lower inputs with the xGen™ RNA Library Prep Kit?
No.
The xGen™ RNA Library Prep Kit does not require adapter titration at any of the supported input levels. Adapter ligation efficiency is maintained at all supported input quantities.
Can I use my own indexing primers with the xGen™ RNA Library Prep Kit?
Go to Appendix H: Indexed adapter sequences in the xGen RNA Library Prep Kit Protocol for details about the adapter sequences and structure of the xGen™ RNA Library Prep Kit adapters and indexing primer kits.
Please contact Scientific Application Support at applicationsupport@idtdna.com if you wish to confirm compatibility of your own custom indexing primers with the stubby adapters supplied in the xGen RNA Library Prep Kit workflows.
Is there a recommended bioinformatics workflow for the xGen™ RNA Library Prep Kit?
Adaptase™ technology, used in the xGen™ RNA Library Prep Kit, adds a low complexity polynucleotide tail with a median length of eight bases to the 3’ end of each fragment during the addition of the second NGS adapter molecule (Read 2 Stubby Adapter). Considering this, it is normal and expected to observe them at the beginning of Read 2 (R2).
When read length is close to fragment size, the tail may also be observed toward the end of Read 1 (R1). We recommend using the STAR aligner (Dobin et al. 2013) as it can soft clip the synthetic tail sequence and provides efficient mapping without additional processing of your sequencing data.
For more details, refer to Appendix G: Data analysis and informatics in the xGen RNA Library Prep Kit Protocol.
Is library amplification required for the xGen™ RNA Library Prep Kit? How many cycles of PCR are needed?
Yes.
The Indexing PCR step is a required step to complete the indexed adapter sequences and generate sufficient library for sequencing.
Refer to the xGen RNA Library Prep Kit Protocol for specific cycle requirements based on input RNA quantity.
Can I generate RNA libraries of different size ranges with the xGen™ RNA Library Prep Kit?
Yes.
Refer to the Appendix section of the xGen RNA Library Prep Kit Protocol titled 'Expected Results with Alternate Fragmentation Times'.
Appendix F provides details on fragmentation times and bead cleanup ratios required for different intended mean library sizes ranging from 380–480 bp (mean insert sizes ranging from 250–350 bp).
Can xGen™ UDI-UMI Adapters be used for RNA-seq?
The xGen™ RNA Library Prep Kit includes a stubby adapter in the workflow so the stocked UDI-UMI adapters are not compatible with those kits.
For customers using homebrew, or other commercial RNA-seq workflows that utilize TA-ligation, these UDI-UMI adapters are compatible. However, it is important to note that stranded RNA-seq libraries constructed using xGen UDI-UMI Adapters have the insert strandedness flipped (i.e., to the opposite strand) when compared to standard Illumina TruSeq™ adapters. As such, this strand flip must be handled during data analysis.
Disclaimer: TruSeq™ and Nextera™ are registered trademarks of Illumina, Inc., used with permission. All rights reserved.


 Processing
Processing